About
The generation of induced pluripotent stem cells (iPSCs), especially the generation of patient-derived pluripotent stem cells suitable for disease modelling in vitro, opens the door for the potential translation of stem-cell related studies into the clinic.
Successful replacement, or augmentation, of the function of damaged cells by patient derived differentiated stem cells would provide a novel cell-based therapy for diseases. Since iPSCs resemble human embryonic stem cells (hESCs) in their ability to generate cells of three germ layers, patient-specific iPSCs offer definitive solutions for the ethical and histo-incompatibility issues related to hESCs. Indeed human iPSC (hiPSC)-based autologous transplantation is heralded as the future of regenerative medicine.
One of our aims is to generate and correct disease-specific hiPSCs for disease modelling and drug screening. The combination of gene-editing based methodologies together with the development of novel protocols for cell differentiation into relevant tissues/organs, provides a unique scenario for modelling disease progression, and the identification of molecular and cellular mechanisms leading to organ regeneration (Figure 1). In this regard we are particularly interested in generation of transgene-free and disease free patient derived hiPSCs for disease modelling and the discovery of novel therapeutic targets.
We believe that the recovery of tissue function should not be restricted to the development of cell replacement therapies. In this regard, in our laboratory we take advantage of organisms that possess the ability to regenerate such as zebrafish, in order to understand which molecular and cellular pathways lead to organ regeneration.
Surprisingly, studies in neonatal mice have demonstrated that soon after birth this organism posses the capability to regenerate its heart. Taking advantage of such preliminary observations we are translating such analysis in order to understand if the mammalian neonatal kidney still posses the capability to regenerate, and more importantly, if we are able to dissect the epigenetic and cellular mechanisms leading to those responses.
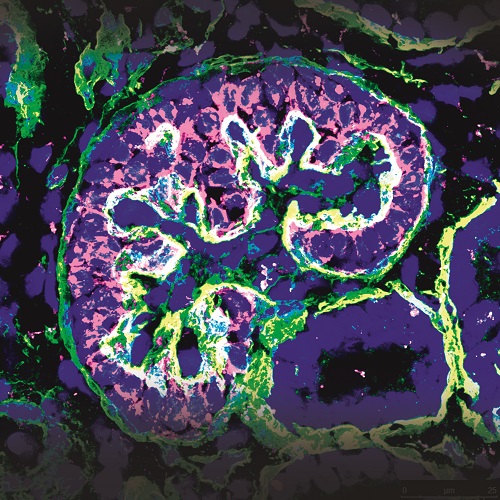
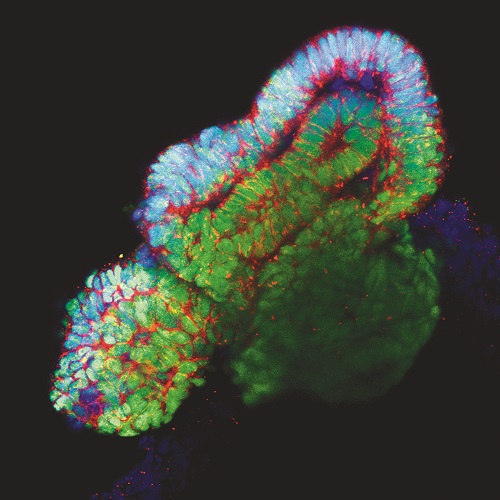
Lastly, and in an effort to fully develop in vitro and ex vivo platforms for organ regeneration, in our lab we are focused in the development of reporter cell lines for different transcription factors essential for tissue-specific commitment and differentiation (i.e: renal and cardiac lineages). The possibility to combine pluripotent stem cell lines together with decellularized matrices, functionalized biomaterials and ex vivo organoids offers and unprecedented opportunity for the immediate generation of patient-specific in vitro and ex vivo platforms for disease modelling and organ regeneration (Figure 2).
Staff
Projects
| NATIONAL PROJECTS | FINANCER | PI |
|---|---|---|
| Bioengeniería para mejorar la salud mediante de organoides y bioimpresión 3D (2021-2023) | MINECO – Plataformas ISCIII de apoyo a la I+D+I en Biomedicina y Ciencias de la Salud | Núria Montserrat |
| CARDIOPRINT Biofabricación avanzada multifunción en 3D para la generación de tejido cardiaco terapéutico a escala humana diseñado por ordenador (2021-2024) | MICIU, Proyectos de I+D+i en líneas estratégicas | Núria Montserrat |
| CAKUTORG Desarrollando nuevas estrategias para entender y tratar las anomalías congénitas del riñón y del tracto urinario mediante organoides (2021-2024) | MICIU, Retos investigación: Proyectos I+D | Núria Montserrat |
| Identifying SARS-CoV-2- host cell interactions exploiting CRISPR/Cas9 engineered human organoids: through the development of specific therapies against COVID19 (2020-2022) | FBBVA | Núria Montserrat |
| CHONDREG · Identification of the epigenetic mechanisms preventing chondrocyte de-differentiation: generation of novel therapeutic strategies for the treatment of cartilage chronic osteochondral lesions | CIBER | Nuria Montserrat |
| Infarto de miocardio en jóvenes. Factores epigeneticos y nuevos marcadores de riesgo cardiovascular. Efecto de la modulación de la expresión de microRNAs y long-non coding RNAs | ISCIII | (Collaborator) |
| INTERNATIONAL FUNDED PROJECTS | FINANCER | PI |
|---|---|---|
| ENGIORG Engineering kidney organoids to study the interplay between Tissue Mechanics and Metabolism: from development to disease (2021-2026) | European Commission | Núria Montserrat |
| ECaBox ECaBox “Eyes in a Care Box”: Regenerating human retina from resuscitated cadaveric eyes (2021-2025) | European Commission, FET OPEN | Núria Montserrat |
| Engineering functional human kidneys and urinary tracts (2021-2024) | Wellcome Leap Solicitation for Humans Organs, Physiology and Engineering (HOPE) | Núria Montserrat |
| BRAV3. Computational biomechanics and bioengineering 3D printing to develop a personalized regenerative biological ventricular assist device to provide lasting functional support to damaged hearts (2020-2024) | European Commission | Núria Montserrat |
| MAD-CoV 2 · Modern approaches for developing antivirals against SARS-CoV 2 (2020-2024) | European Commission | Núria Montserrat |
| R2U-Tox-Assay · Ready-to-use Toxicity Screening Assay based on iPS-Technologies (2020-2022) | EIT Health | Núria Montserrat |
| PRIVATELY FUNDED PROJECTS | FINANCER | PI |
|---|---|---|
| SYSTORG Exploiting organoid model systems to explore systemic conditions worsening COVID19: merging cellular and genetic engineering (2021-2024) | Fundació La Marató de TV3, TV3-Projectes de recerca La Marató TV3 | Núria Montserrat |
| Identificació de noves dianes terapèutiques i biomarcadors de progressió del càncer de ronyó a través de models organoides i xenoempelts genèticament dissenyats per CRISPR (2020-2023) | Fundació La Marató de TV3, TV3-Projectes de recerca La Marató TV3 | Núria Montserrat |
| REPIRE · Regenerating photoreceptors in human retinal organoids to establish a treatment for Retinitis Pigmentosa (2018-2021) | Fundación Bancaria “La Caixa” | Núria Montserrat |
| FUNDRAISING PROJECTS | FINANCER | PI |
|---|---|---|
| Programa Faster Future 2020: COVID-19 (2021) | Fundraising | Núria Montserrat |
| FINISHED PROJECTS | FINANCER | PI |
|---|---|---|
| Identification of Kidney Cancer progression targets and biomarkers through CRISPR-engineered organoids and xenograft mouse models (2019-2020) | Fundació La Marató de TV3 | Núria Montserrat |
| Generation of Isogenic Models of Clear Cell Renal Cell Carcinoma (ccRCC) using CRISPR-engineered Kidney Organoids, for the identification of diagnostic biomarkers (2017-2021) | Fundación AECC | Núria Montserrat |
| EPIORG · Cómo modelar la Nefropatía Diabética: restableciendo el epigenoma en organoides renales diabéticos inducidos (2018-2020) | MINECO, Retos investigación: Proyectos I+D | Núria Montserrat |
| MECHANORG · Como integrar señales mecánicas y metabólicas en organoides renales para el modelado de patologías humanas (2019-2020) | MINECO, Acciones Dinamización Europa Investigación | Núria Montserrat |
| Modelling Diabetic Nephropathy targeting DNA methylation: engineering the epigenome in kidney (2019-2020) | EFSD European Foundation for the Study of Diabetes | Núria Montserrat |
| REGMAMKID · How to regenerate the mammalian kidney (2015-2021) | European Commission, ERC-StG | Núria Montserrat |
| REPROMICRO · Reprogramacion y regeneracion tisular a partir de microvesiculas derivadas de celulas madre de pluripotencia inducida (2017-2019) | Ministerio de Economía y Competitividad, Explora Ciencia | Nuria Montserrat |
| Desarrollo de nuevas estrategias para el tratamiento de la enfermedad renal (2015-2017) | MINECO | Nuria Montserrat |
| TRATENFREN · Desarrollo de nuevas estrategias para el tratamiento de la enfermedad renal (2015-2017) | MINECO, Retos investigación: Proyectos I+D | Nuria Montserrat |
| Regenerative medicine for Fanconi anemia: generation of disease-free patient-specific iPS (2013-2016) | Fundació La Marató de TV3 | Nuria Montserrat |
| ACE2-ORG · Development of a human cellular plaform unveilling Angiotensin-converting enzyme 2 (ACE2) – sars-CoV-2 interactions (2020-2021) | ISCIII | Núria Montserrat |
| Red TERCEL · Red de Terapia Celular (2017-2021) | MINECO, ISCIII | (Collaborator) |
| EPIORGABOLISM Diabetic nephropathy modelling in hESC-derived 3D (2019-2021) | European Commission, MARIE CURIE – IF | Carmen Hurtado |
Publications
Check for more detailed information on the outputs of the Group at IBEC CRIS portal.
Publications list:
Equipment
- Real Time QuantStudio 5
- SimpliAmp thermocycler
- Eppendorf 5415D centrifuge
- Allegra X-15 R centrifuge
- Gyrozen 1248 centrifuge
- BioUltra 6 Telstar culture Hood 2x
- AH-100 Telstar primary culture Hood
- Binder CB 60 incubators 2x
- Controltecnica ASTEC SCA 165 incubator
- Controltecnica ZC 180 incubator
- Bioruptor Pico sonicator
- Thermomixer C thermal block
- Leica DMS1000 and DMIL Led microscopes
- Leica DMi1 microscope
- Leica MZ 10F magnifying glass
- Safe Imager 2.0 transilluminator
Collaborations
- Juan Carlos Izpisua Belmonte
Salk Institute for Biological Studies - Dr. Josep Maria Campistol Plana
Experimental Laboratory of Nephrology and Transplantation, Hospital Clínic, Barcelona - Peter Hohestein
The Roslin Institute, University of Edinburgh - Dr. Pere Gascón Vilaplana
Head of Oncology Service/Molecular and Translational Oncology Laboratory, IDIBAPS - Gloria Calderon
Embryotools SL - Pura Muñoz Cánovas
Departament de Ciències Experimentals i de la Salut, Universitat Pompeu Fabra - Dr. Pedro Guillén
Director Clínica Cemtro, Madrid - Dr. Francisco Fernández Avilés
Head of Cardiology Service, Hospital General Universitario Gregorio Marañón, Madrid - Dr María Eugenia Fernández
Unit of Cell Production, Hospital Gregorio Marañón, Madrid - Joaquin Gutiérrez Fruitós
University of Barcelona - Dr. Pere Roca-Cusachs
IBEC - Dr. Elena Martínez
IBEC - Dr. Cristina Eguizabal Argaiz
Centro Vasco de Transfusion y Tejidos Humanos (CVTTH), Bizkaia - Dr. Antonio Alcaraz
Head of Urology, Hospital Clínic, Barcelona - Dr. Oriol Casanovas
Head of Tumour Angiogenesis Group, IDIBELL
News
La Vanguardia: Los organoides de riñón producidos en serie abren la vía a mejorar los trasplantes
Los bioingenieros descubren cómo obtener los tejidos de manera rápida y asequible.
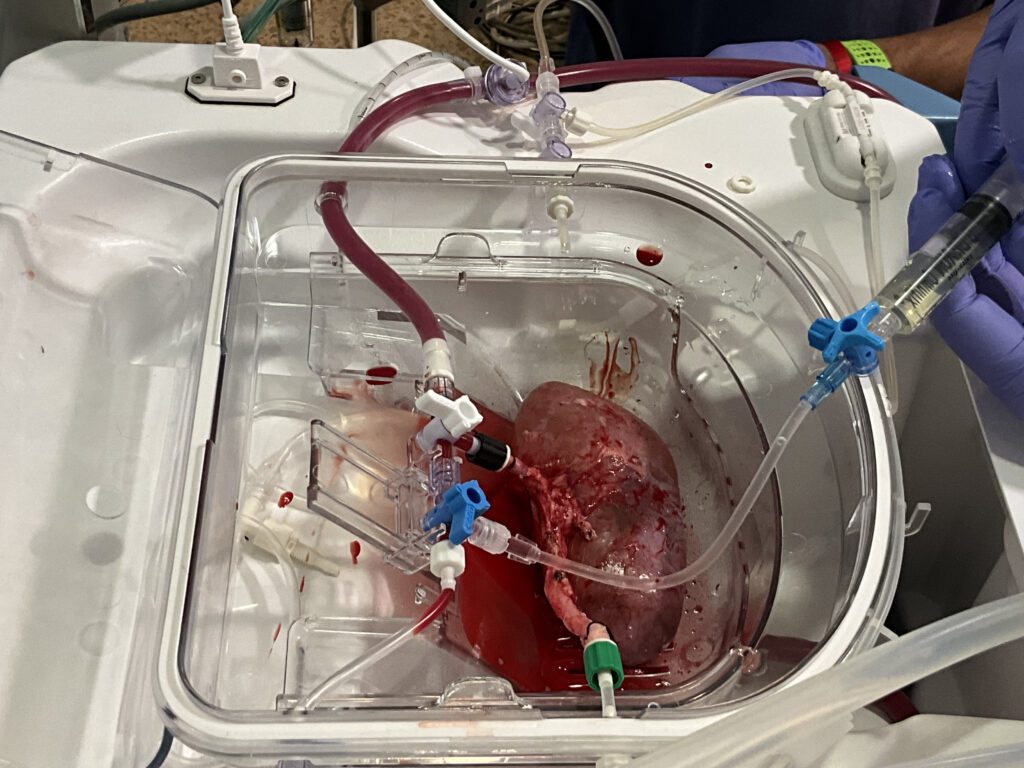
EL MUNDO: Prueban con éxito organoides humanos para reparar riñones
Científicos españoles demuestran la viabilidad de una tecnología celular con la que abordar las complicaciones en los trasplantes. El experimento ha demostrado que “la producción es escalable y segura”.
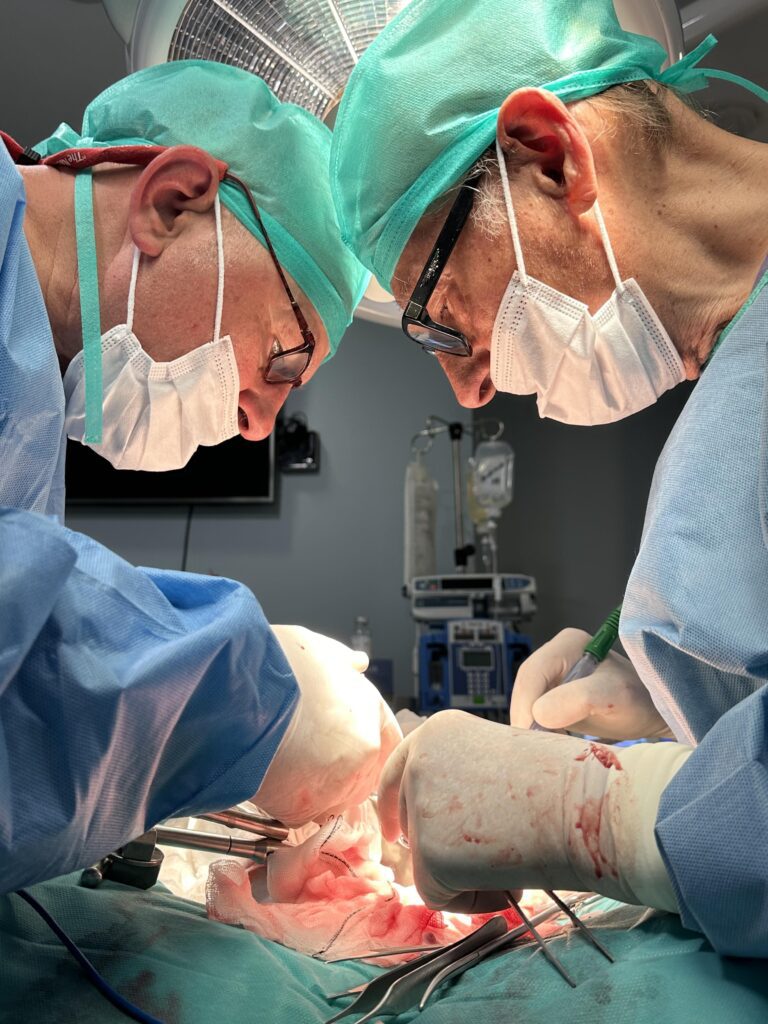
La Razón: Trasplantan a cerdos riñones modificados con organoides humanos
El objetivo es reparar o regenerar un órgano antes del implante para disponer de más que sean viables.
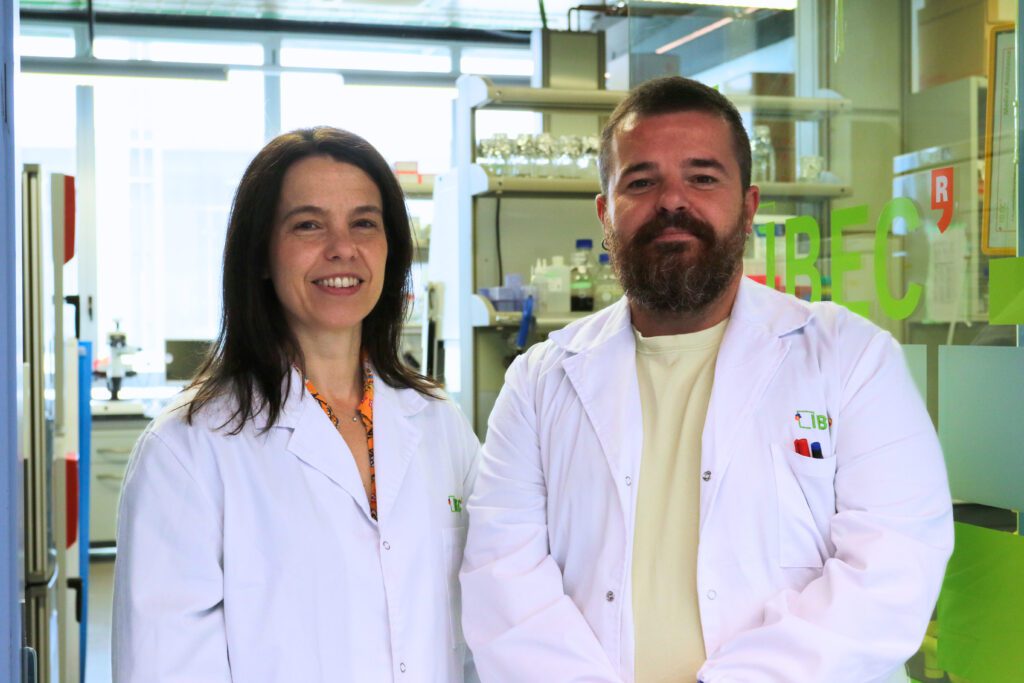

First transplant in pigs of modified porcine kidneys with human renal organoids
A research team has developed pioneering technology that enables human kidney organoids to be produced on a scalable basis. These organoids can then be combined with pig kidneys outside the body and transplanted back into the same animal in a viable manner. The experiment, led by the Institute for Bioengineering of Catalonia (IBEC), is in the preclinical phase. It confirms the safety and viability of the procedure, paving the way for future trials involving humans. In the long term, this approach could help to extend the useful life of organs intended for transplantation and provide an alternative therapy for patients with chronic kidney disease.
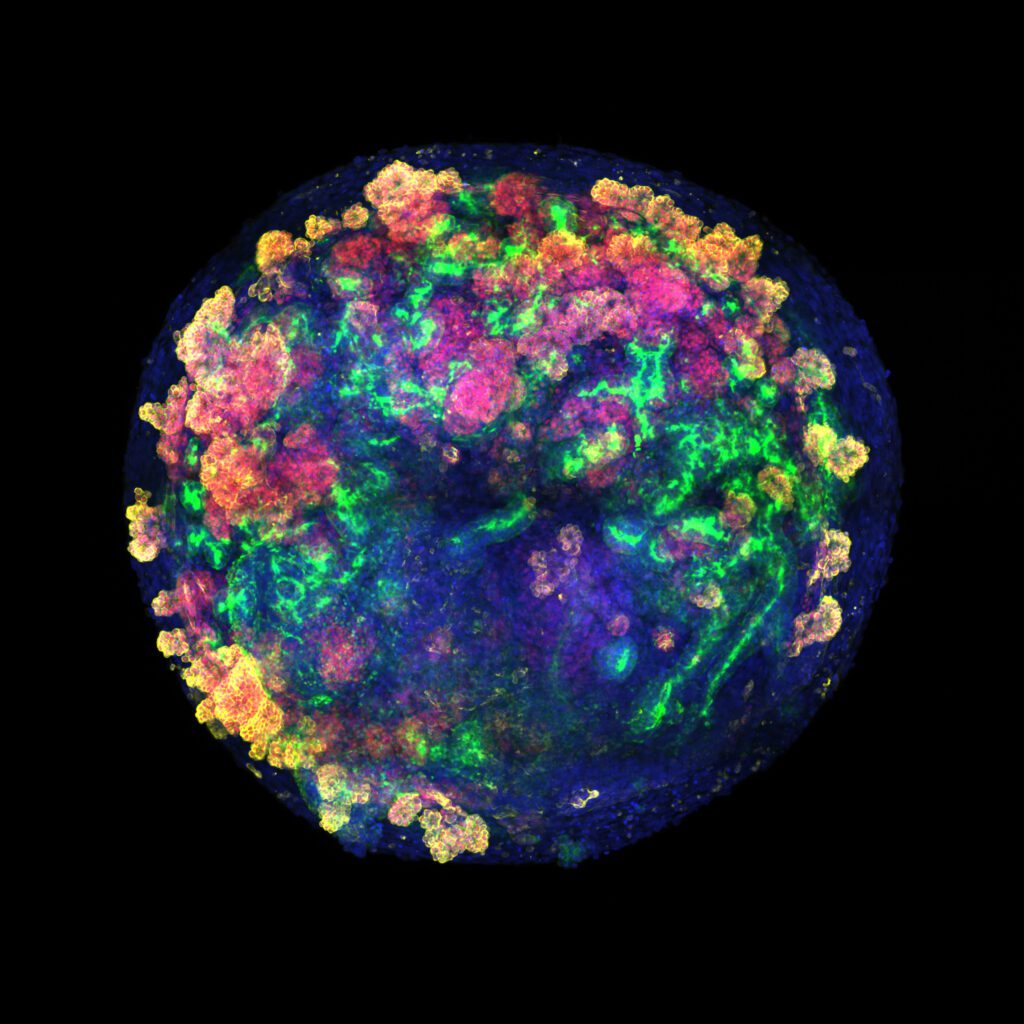
El País: El primer trasplante de minirriñones humanos en cerdos abre una vía para no desperdiciar miles de órganos donados
El avance puede facilitar la recuperación de vísceras dañadas para reducir las listas de espera de trasplantes, que no dejan de crecer
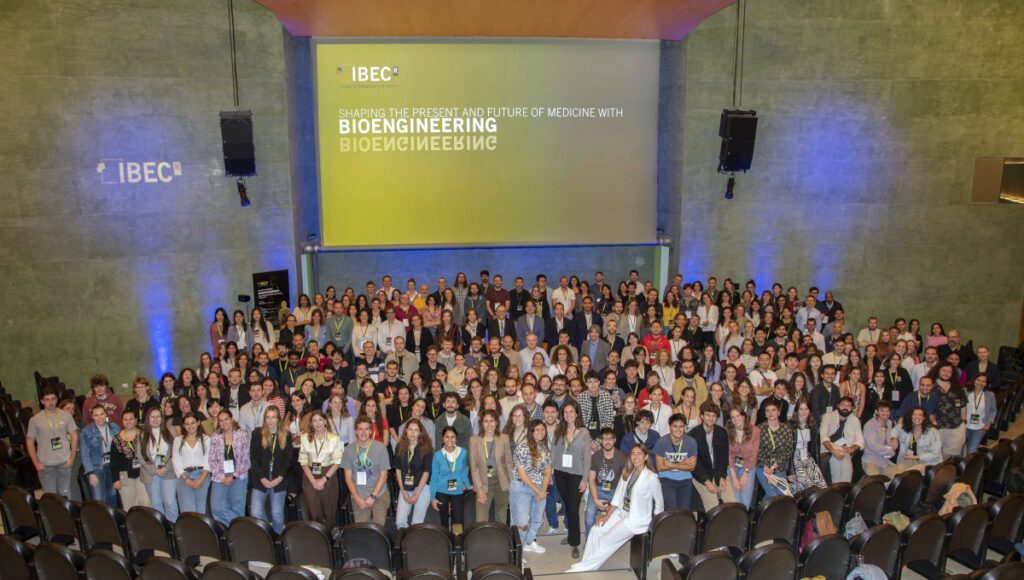

Bioengineering for Emergent and Advanced Therapies at the 17th IBEC Symposium
IBEC’s 17th Annual Symposium focused on ‘Bioengineering for Emergent and Advanced Therapies’, one of IBEC’s key application areas. Around 300 people attended the event, including local and international researchers. It was a multidisciplinary environment in which experts from other centres and the IBEC community itself had the opportunity to present their projects and share knowledge.

Núria Montserrat receives funding from Aefat to advance research on Ataxia Telangiectasia
Núria Montserrat, ICREA Research Professor at the Institute for Bioengineering of Catalonia (IBEC), has been awarded two significant grants totaling 50,000 euros. These funds are intended to support an innovative … Read more
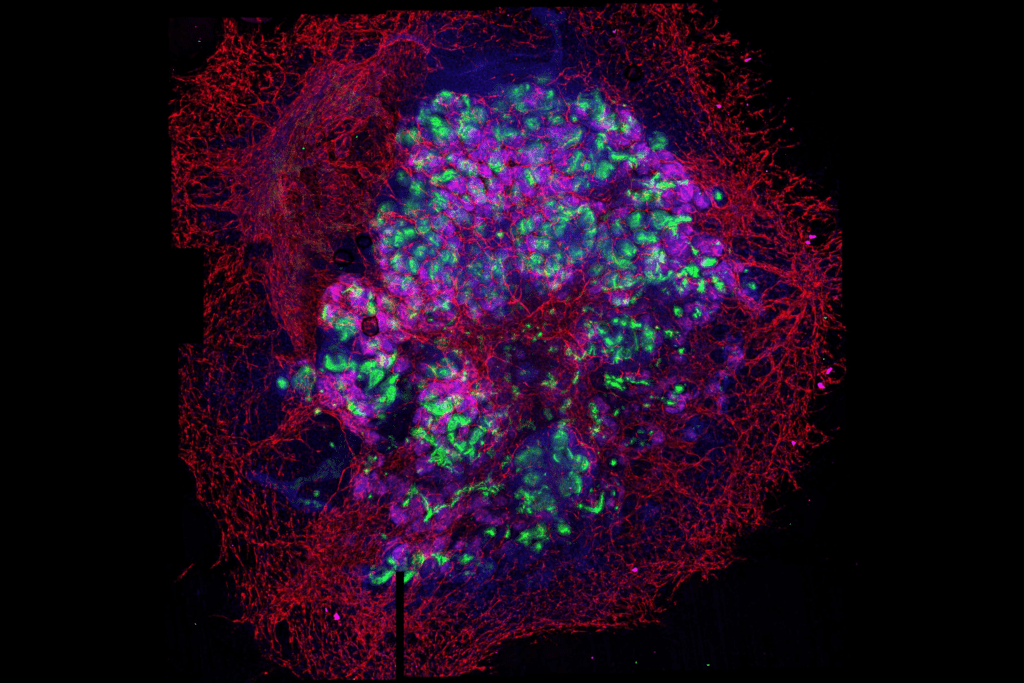
For the first time scientists managed to generate kidney organoids with a complex vascular system
These are asembloids, micrometric organoids formed by the union of kidney organoids with vascular organoids in 3D. These small culture systems are useful for disease modelling and drug screening. The study, led by IBEC, describes an approach that has not been tried before.
Jobs
Postdoctoral researcher at the at the Pluripotency for organ regeneration Research Group
Ref: PD_EG //Deadline: 31/10/2024
PhD student at the Pluripotency for organ regeneration Research Group
Ref: PhD-NM//Deadline:16/12/2023
Postdoc at the Pluripotency for organ regeneration Research Group (ENGINORG)
Ref: PD-NM//Deadline: 15/12/2023
Laboratory Technician at the Pluripotency for organ regeneration Research Group (LT-NM)
Ref.: LT-NM // Deadline: March 6th
Postdoc at the Pluripotency for organ regeneration Research Group (ENGINORG)
Ref.: PD-NM / Deadline: 30/01/2023
Postdoc at the Pluripotency for organ regeneration Research Group (REACT)
Ref.: PD-NM / Deadline: 15/12/2023
Senior Laboratory Technician at the Pluripotency for Organoid Regeneration Research Group (SLT_NM)
Ref: SLT_NM // Deadline: 16/01/2023
Laboratory Technician at the Pluripotency for Organ Regeneration Research Group (LT_NM)
Ref: LT_NM // Deadline: 16/01/23
Junior Project Manager at the Plataforma ISCIII Biobancos y Biomodelos (Ref.: PM2-NM)
Ref: PM2-NM // Deadline: 18/03/2024